Forecasting CWD with Physics-Informed GNNs
Spatio-temporal GNN (GATv2 + GRU) with Metapopulation-SIR hybrid loss for chronic wasting disease forecasting across 1,438 US counties.
Read paper →Chronic wasting disease is a fatal prion disease in deer, elk, and moose with no known treatment. Surveillance data is sparse, zero-inflated, and spatially heterogeneous (most county-years report zero cases because no one tested) which breaks both naive regression and pure mechanistic SIR modeling.
Our team built a county graph (1,438 nodes, 8,064 edges) from a 31k-record surveillance dataset across 16 U.S. states, then trained a spatio-temporal GNN with a physics-informed loss that combines data fit with a one-step Metapopulation-SIR residual.
Architecture
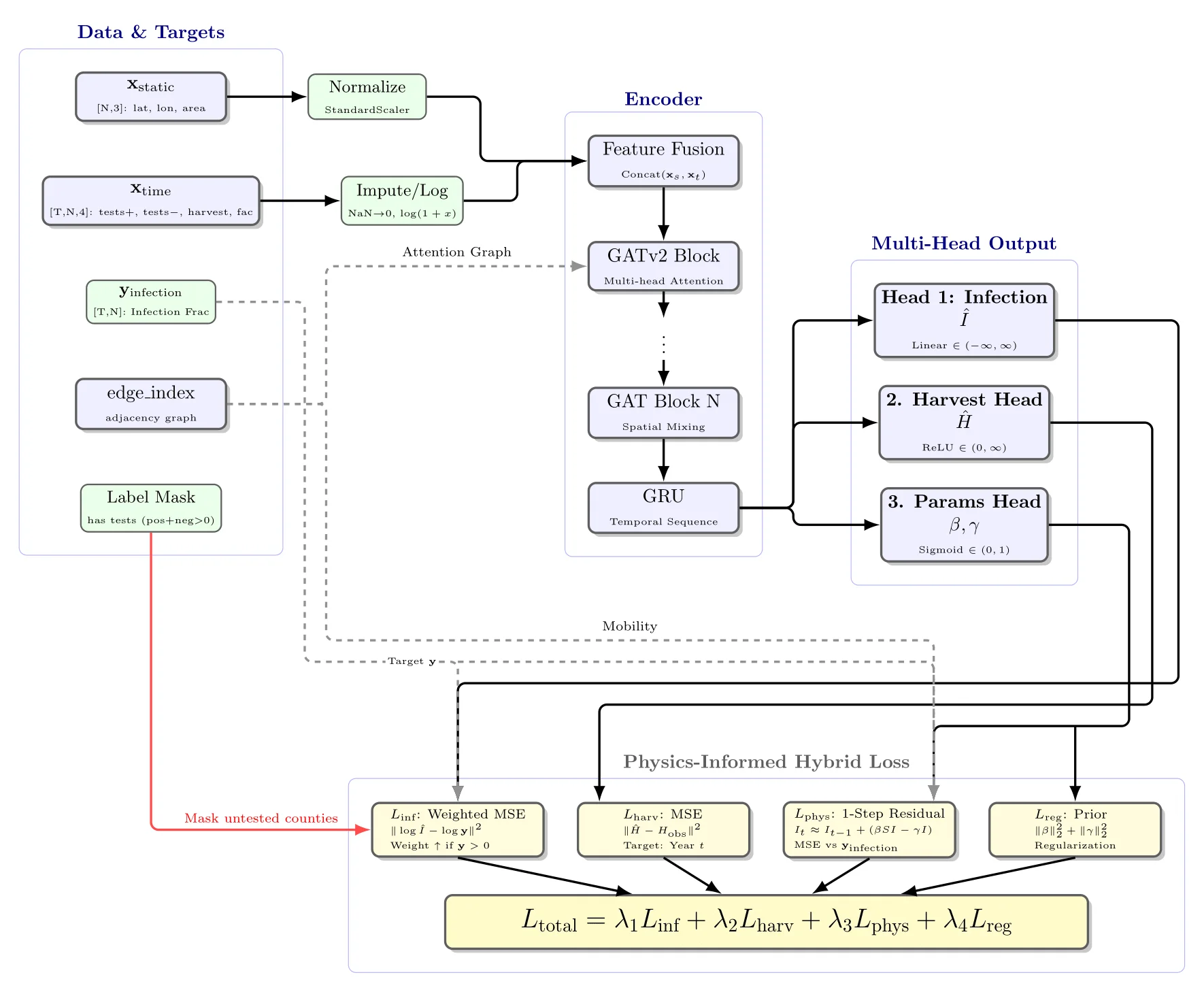
The encoder fuses static features (lat/lon/area) and time-varying features (positive/negative tests, harvest, facilities) through GATv2 attention blocks and a GRU. Three heads decode infections, harvests, and the per-county SIR parameters . The physics term enforces that consecutive infection predictions roughly satisfy the SIR update regularizing the GNN toward epidemiologically plausible trajectories without forcing a fixed ODE form.
Forecast — 2020–2021 season
| Ground truth | Predicted (GNN + harvest + SIR loss) |
|---|---|
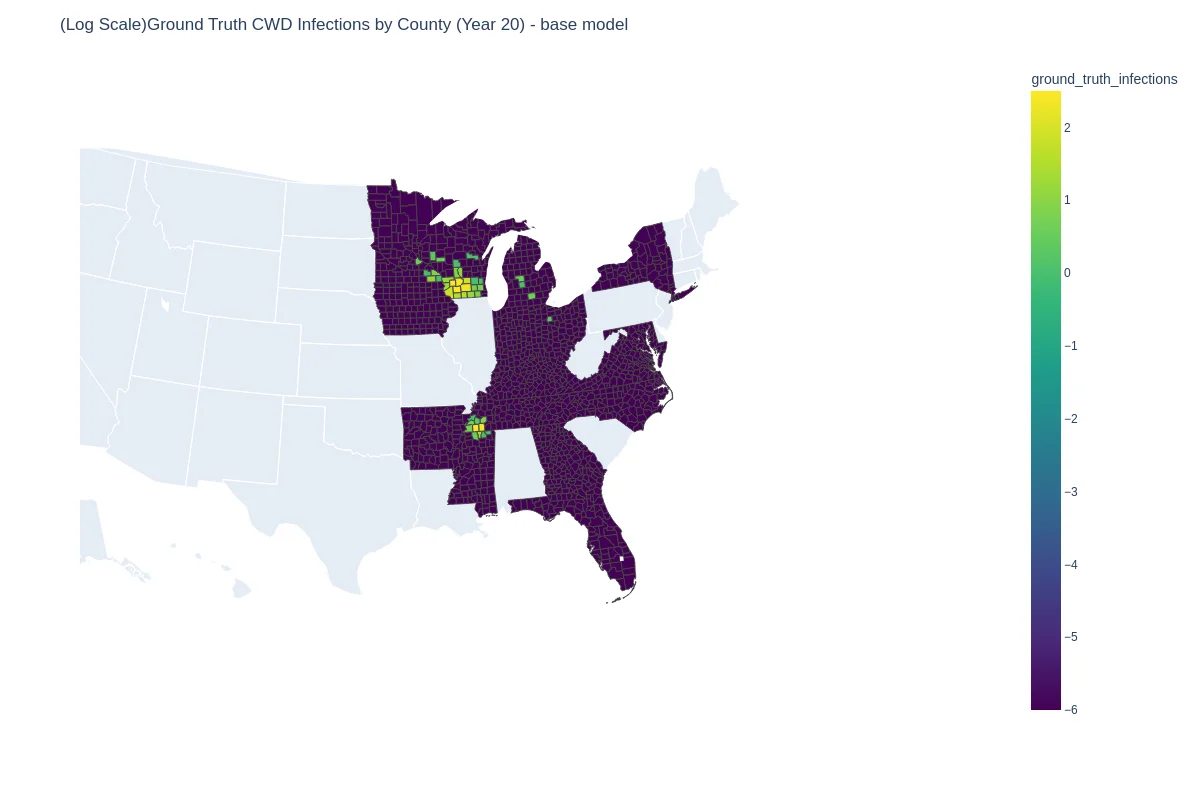 | 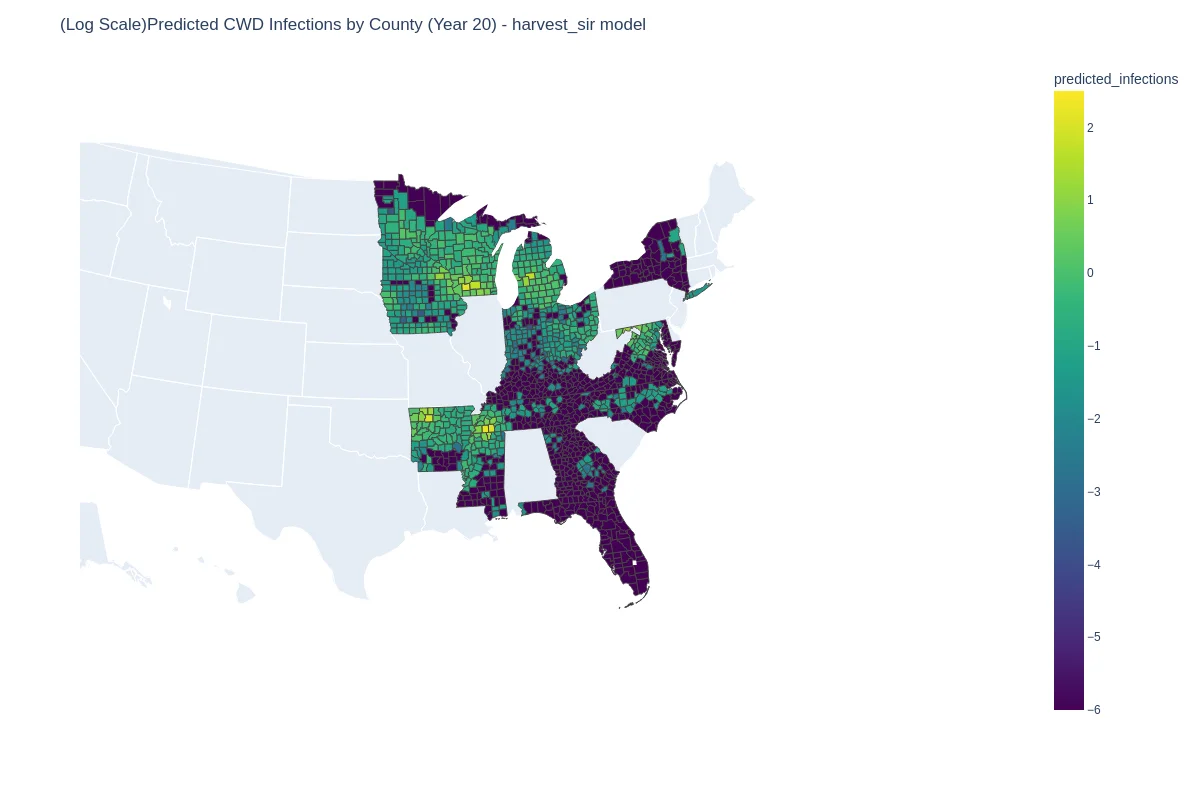 |
The full hybrid model hit on held-out 2020–2021 infections, outperforming (1.08) and an ODE-only Metapopulation-SIR baseline (2.62), while keeping interpretable SIR parameters at the county level.